CAM-UV
Model
This method CAM-UV (Causal Additive Models with Unobserved Variables) [2] assumes an extension of the basic CAM model [1] to include unobserved variables. This method makes the following assumptions:
The effects of direct causes on a variable form generalized additive models (GAMs).
The causal structures form directed acyclic graphs (DAGs).
CAM-UV allows the existence of unobserved variables. It outputs a causal graph where a undirected edge indicates the pair of variables that have an unobserved causal path (UCP) or an unobserved backdoor path (UBP), and a directed edge indicates the causal direction of a pair of variables that do not have an UCP or UBP.
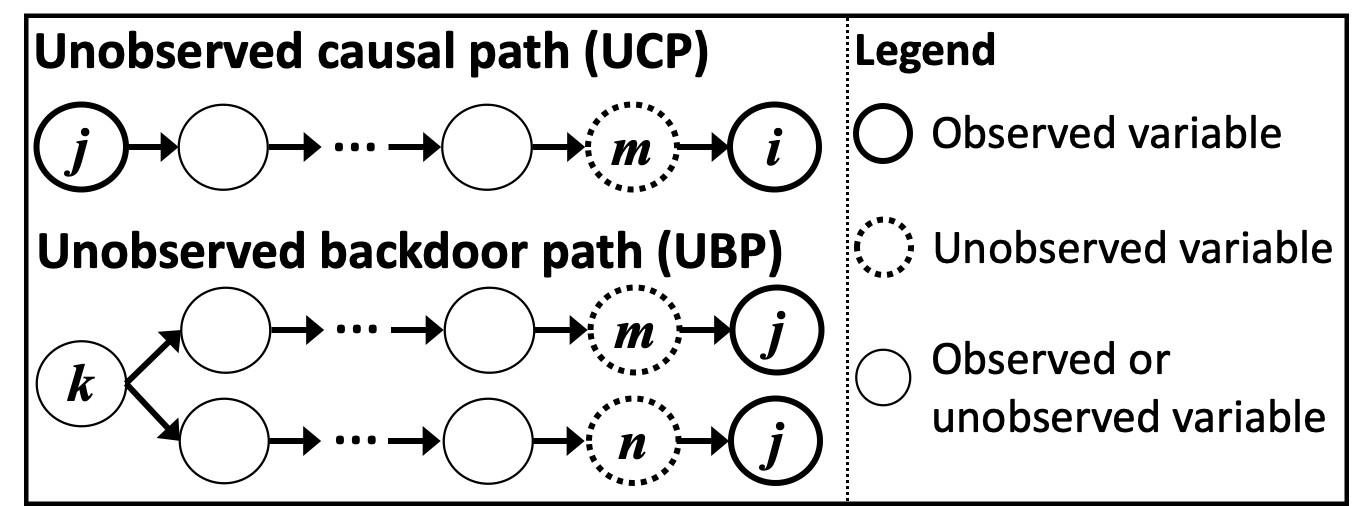
Definition of UCPs ans UBPs: As shown in the below figure, a causal path from \(x_j\) to \(x_i\) is called an UCP if it ends with the directed edge connecting \(x_i\) and its unobserved direct cause. A backdoor path between \(x_i\) and \(x_j\) is called an UBP if it starts with the edge connecting \(x_i\) and its unobserved direct cause, and ends with the edge connecting \(x_j\) and its unobserved direct cause.
References
Import and settings
In this example, we need to import numpy in addition to lingam.
import numpy as np
import random
import lingam
Test data
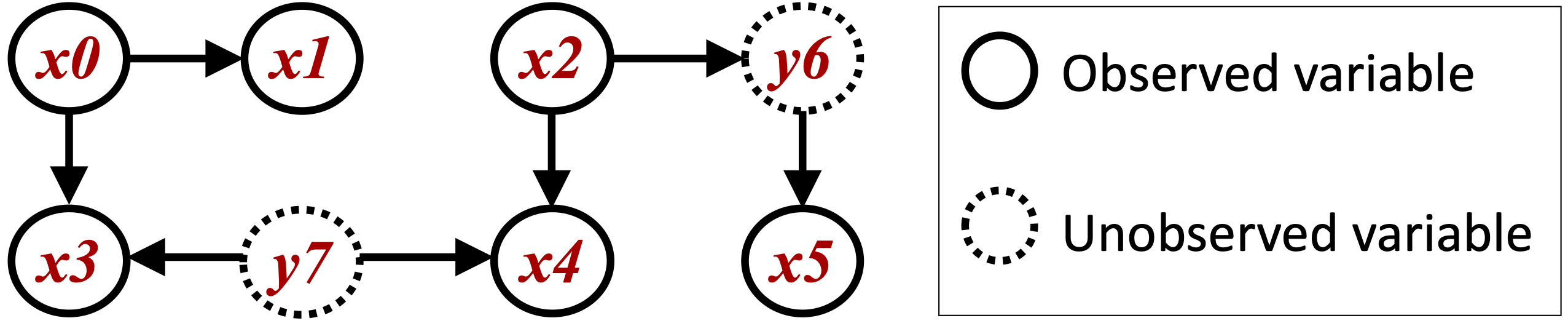
First, we generate a causal structure with 2 unobserved variables (y6 and y7) and 6 observed variables (x0–x5) as shown in the below figure.
def get_noise(n):
noise = ((np.random.rand(1, n)-0.5)*5).reshape(n)
mean = get_random_constant(0.0,2.0)
noise += mean
return noise
def causal_func(cause):
a = get_random_constant(-5.0,5.0)
b = get_random_constant(-1.0,1.0)
c = int(random.uniform(2,3))
return ((cause+a)**(c))+b
def get_random_constant(s,b):
constant = random.uniform(-1.0, 1.0)
if constant>0:
constant = random.uniform(s, b)
else:
constant = random.uniform(-b, -s)
return constant
def create_data(n):
causal_pairs = [[0,1],[0,3],[2,4]]
intermediate_pairs = [[2,5]]
confounder_pairs = [[3,4]]
n_variables = 6
data = np.zeros((n, n_variables)) # observed data
confounders = np.zeros((n, len(confounder_pairs))) # data of unobserced common causes
# Adding external effects
for i in range(n_variables):
data[:,i] = get_noise(n)
for i in range(len(confounder_pairs)):
confounders[:,i] = get_noise(n)
confounders[:,i] = confounders[:,i] / np.std(confounders[:,i])
# Adding the effects of unobserved common causes
for i, cpair in enumerate(confounder_pairs):
cpair = list(cpair)
cpair.sort()
data[:,cpair[0]] += causal_func(confounders[:,i])
data[:,cpair[1]] += causal_func(confounders[:,i])
for i1 in range(n_variables)[0:n_variables]:
data[:,i1] = data[:,i1] / np.std(data[:,i1])
for i2 in range(n_variables)[i1+1:n_variables+1]:
# Adding direct effects between observed variables
if [i1, i2] in causal_pairs:
data[:,i2] += causal_func(data[:,i1])
# Adding undirected effects between observed variables mediated through unobserved variables
if [i1, i2] in intermediate_pairs:
interm = causal_func(data[:,i1])+get_noise(n)
interm = interm / np.std(interm)
data[:,i2] += causal_func(interm)
return data
sample_size = 2000
X = create_data(sample_size)
Causal Discovery
To run causal discovery, we create a CAMUV object and call the fit
method.
model = lingam.CAMUV()
model.fit(X)
Using the adjacency_matrix_ properties, we can see the adjacency matrix as a result of the causal discovery. When the value of a variable pair is np.nan, the variables have a UCP or UBP.
model.adjacency_matrix_
array([[ 0., 0., 0., 0., 0., 0.],
[ 1., 0., 0., 0., 0., 0.],
[ 0., 0., 0., 0., 0., nan],
[ 1., 0., 0., 0., nan, 0.],
[ 0., 0., 1., nan, 0., 0.],
[ 0., 0., nan, 0., 0., 0.]])